 For sequencing itself to occur, the lane mix of prepared libraries is strongly diluted following library preparation.įor example, a common scenario would involve diluting a lane mix down to 2 nM, followed by a second dilution by a factor of around 1,000 to a concentration of 2 pM which is used to load on the sequencer. One way to look at this is by working backwards, starting with the needs of the sequencer – more specifically, the flow cell. One key facet of sequencing lies in the relationship between the amplification and QC of the libraries, and the sequencing run itself. After completion of the sequencing run, the optical signals are translated into the actual sequence, a process termed base calling (Fig. The termination block and the nucleotides of the previous cycle are removed, and the process is repeated until the desired read length is reached. 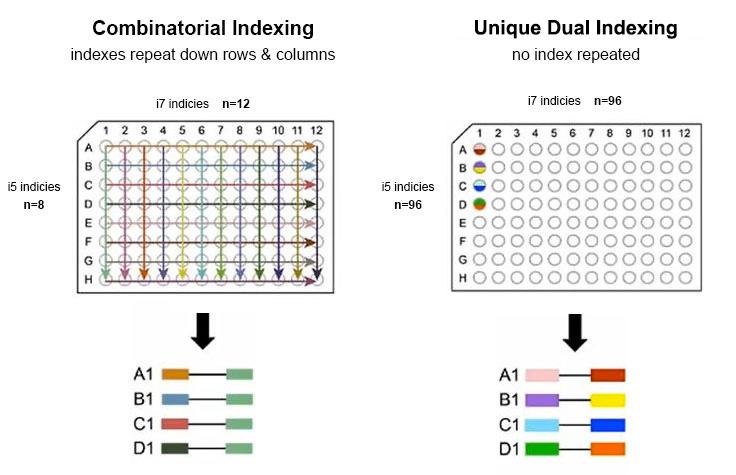
After each cycle, the emission of the fluorophore is detected for each cluster using an optical system and thus the identity of the incorporated base is determined. Figure 2 illustrates the process with four fluorescently labeled nucleotides (Courtesy of Illumina, Inc.). The label also acts as a terminator, so that the reaction is stopped after the incorporation of one nucleotide. Depending on the chemistry used in the respective machine, either all four nucleotides are labelled differently, or only a subset of the nucleotides contain a label. This ensures that only the forward strands are sequenced and the read out is homogenous and not overlayed with a second sequence which would otherwise render the detected sequence unusable.Ī polymerase then “sequences” the insert of the library by adding nucleotides to the complementary strand that are fluorescently labelled. Essentially, enough material needs to be generated by clonal amplification so that the signals generated in the sequencing process become detectable.įirst, reverse strands are removed before sequencing starts. After forward and reverse strands are generated and both are stably attached to the flow cell, clusters are generated by clonal amplification, i.e., the process is repeated over and over until the required level of amplification is reached. This second oligo is complementary to the other sequencing adapter, and thus upon elongation the reverse strand is generated. The original molecule is washed away, and the strand bends over (like a “bridge”) to attach to the next flow cell oligo. Thereby, a copy of the molecule to be sequenced is now covalently attached to the flow cell. 
During this process, the complementary strand is generated by elongating the oligo attached to the flow cell. Once attached, clusters can be generated, and libraries are sequenced by synthesis.Ĭluster generation begins by bridge amplification. The complementary oligonucleotides on the flow cell surface act as an anchor to capture the libraries by binding to the outer region of the Illumina adapter sequence. In the first step, the single-stranded sequencing-ready libraries are loaded onto the flow cell. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
